The University of ALabama at Birmingham CANcer data analysis Portal
Please cite: Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Rodriguez IP, Chakravarthi BVSK and Varambally S. UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017 Aug;19(8):649-658. doi: 10.1016/j.neo.2017.05.002 [PMID:28732212]
Step 1: Go to analysis page of UALCAN and choose cancer of interest from left panel by clicking it.
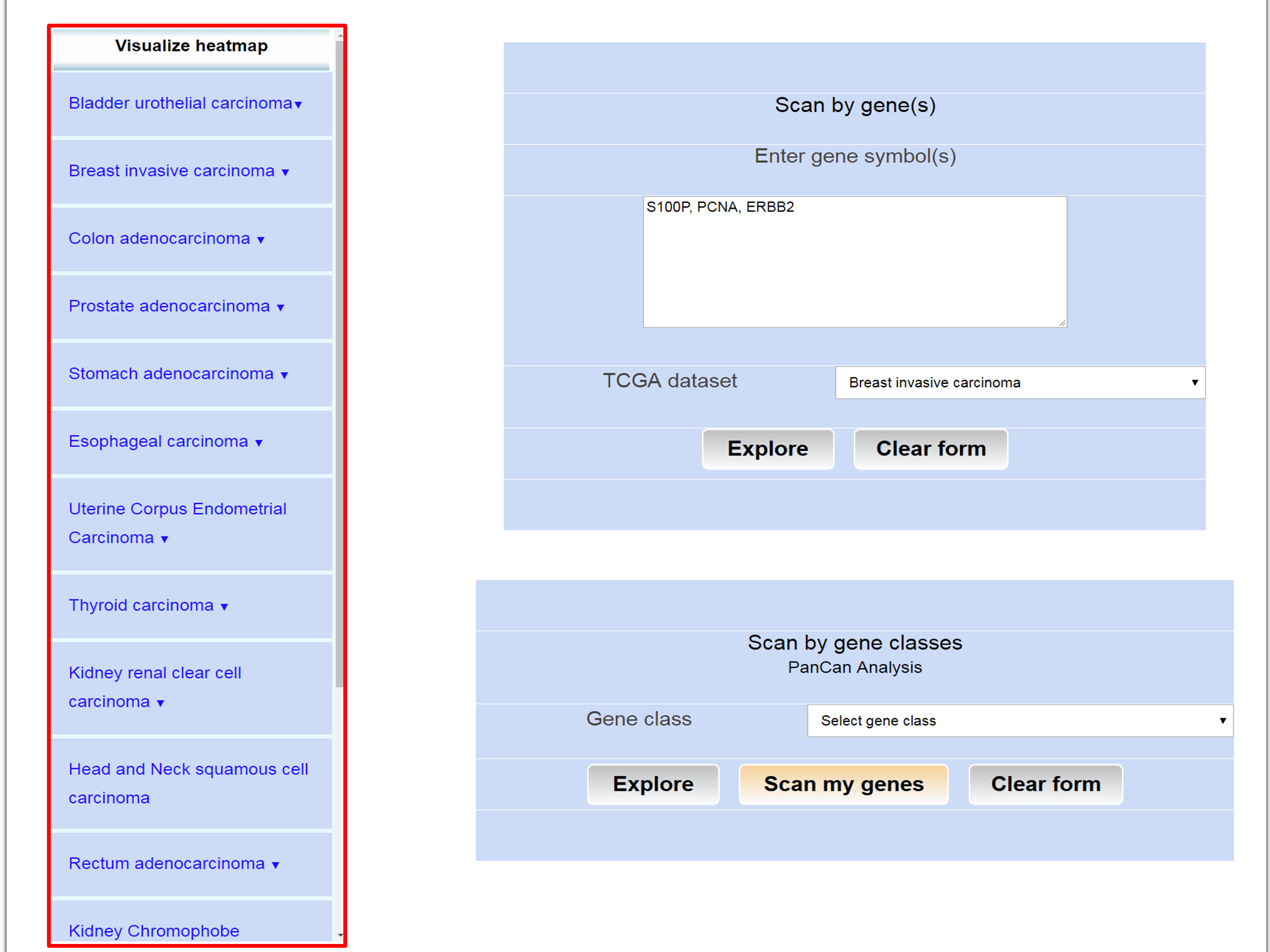
Step 2: UALCAN lists top 250 over-/under-expressed genes in cancer of interest compared to normal samples.
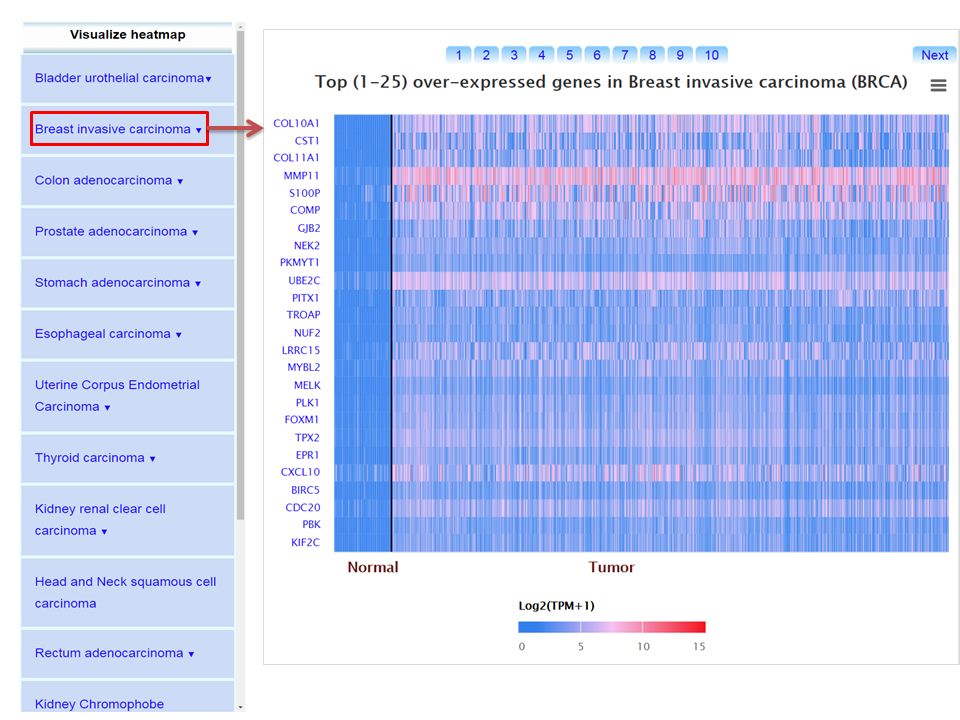
Step 3: User can analyze gene expression and survival profiles of individual gene by clicking the on the heat map.
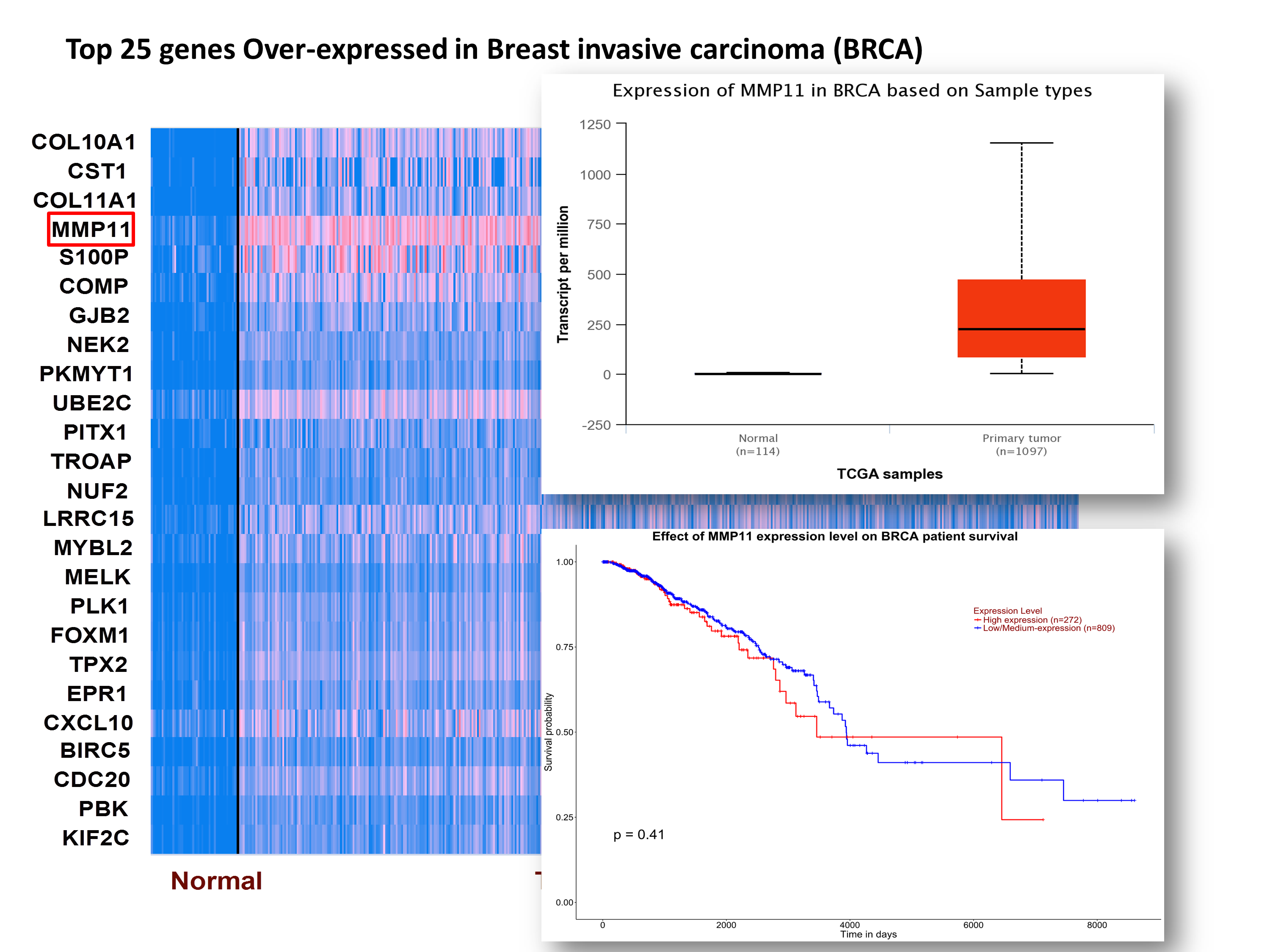
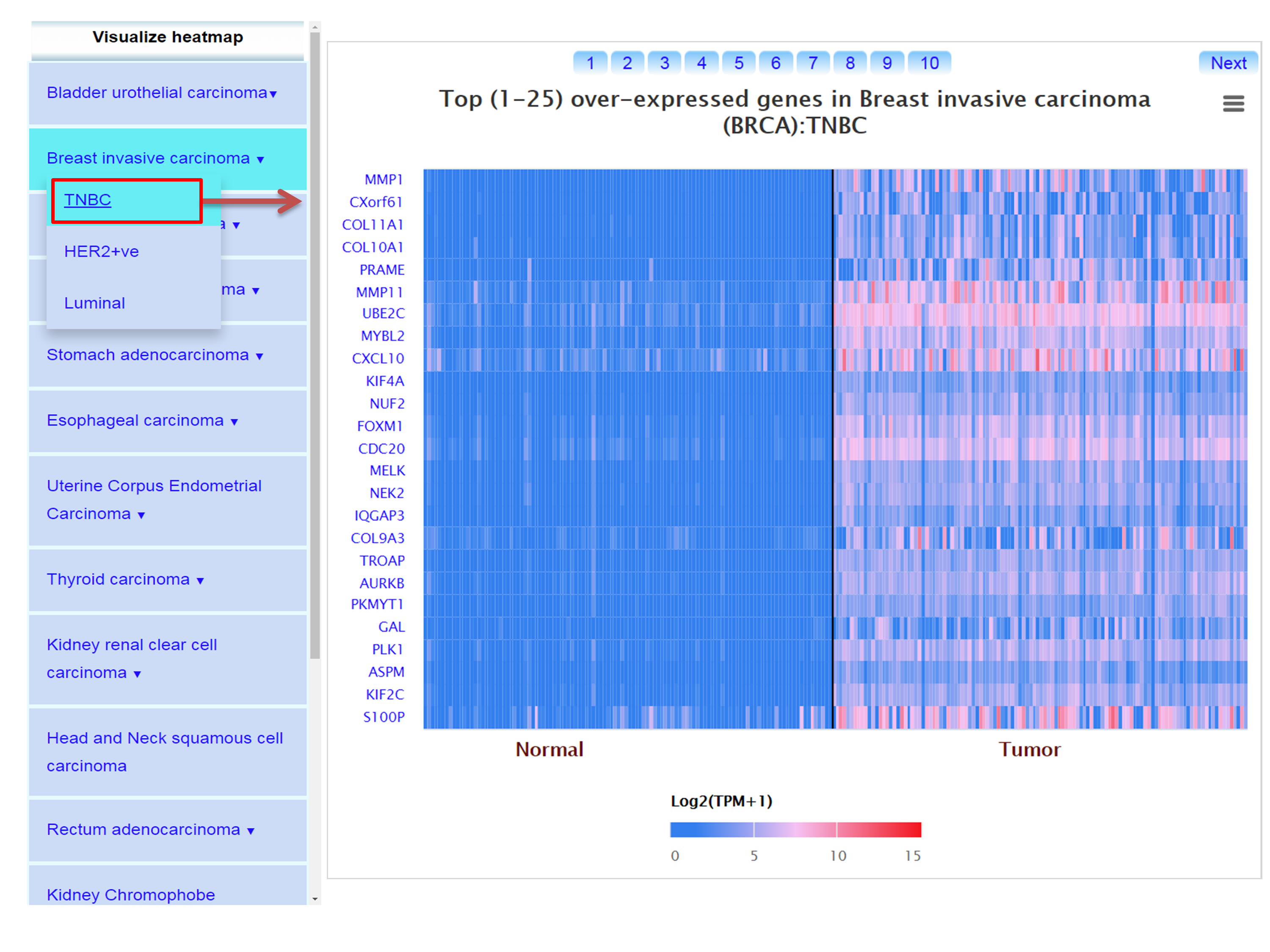
Step 4: UALCAN also enables user to view top 250 over-/under-expressed genes in major cancer subtypes compared to normal samples.
Step 1: Go to analysis page of UALCAN and enter offical symbol of gene(s) in the text area
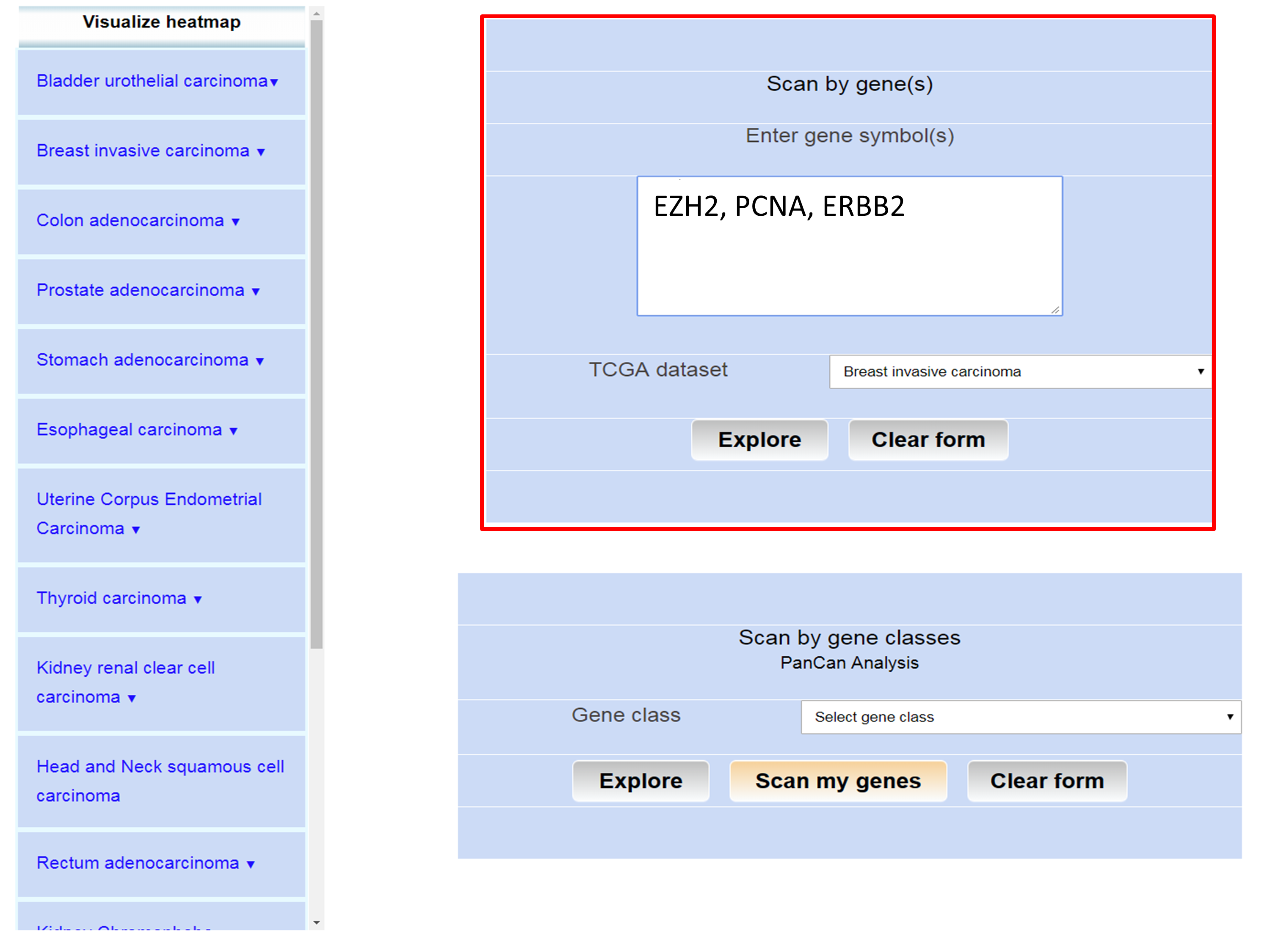
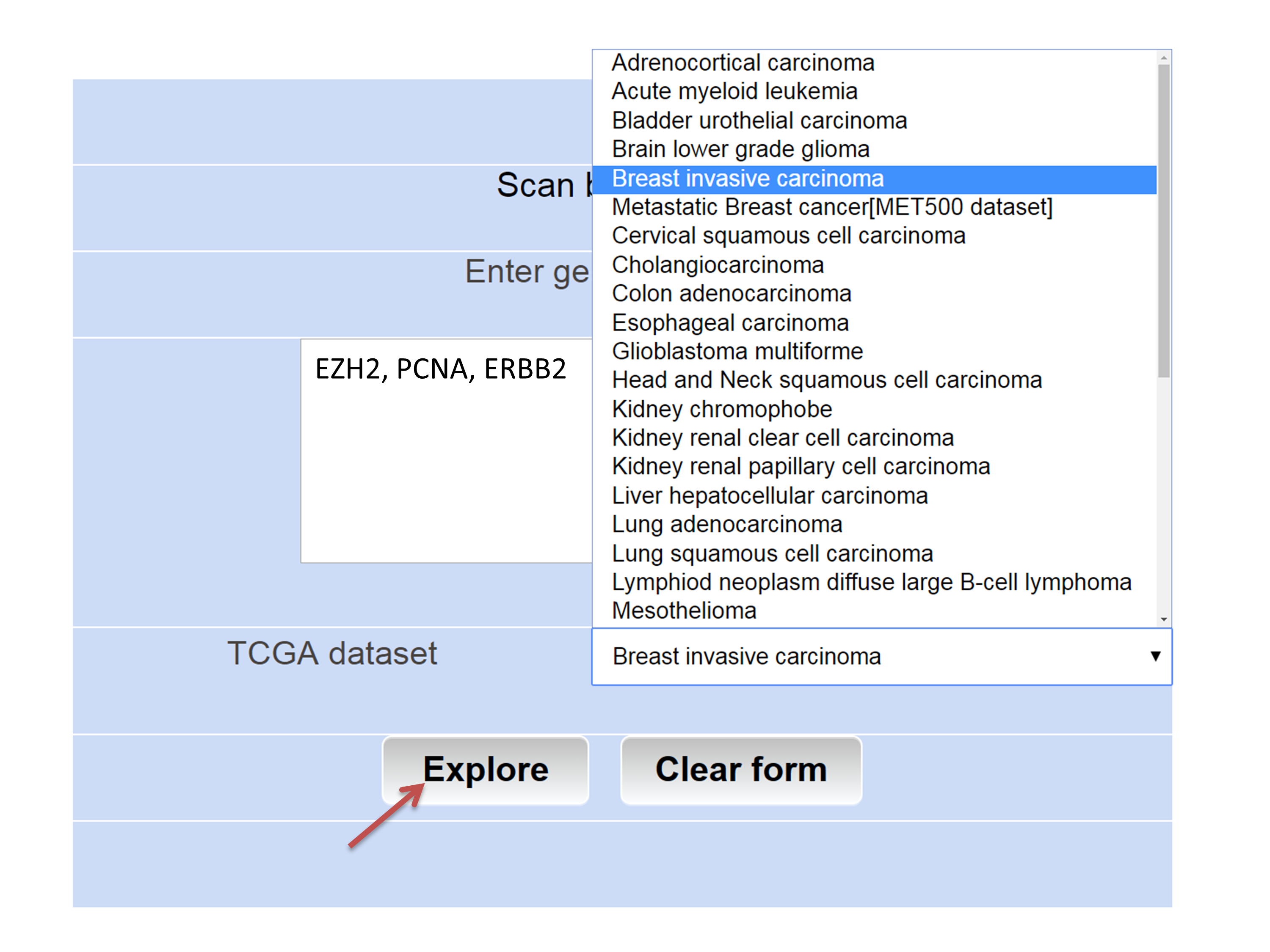
Step 2: Choose the TCGA data set of interest from the drop down menu and click "Explore" button to submit.

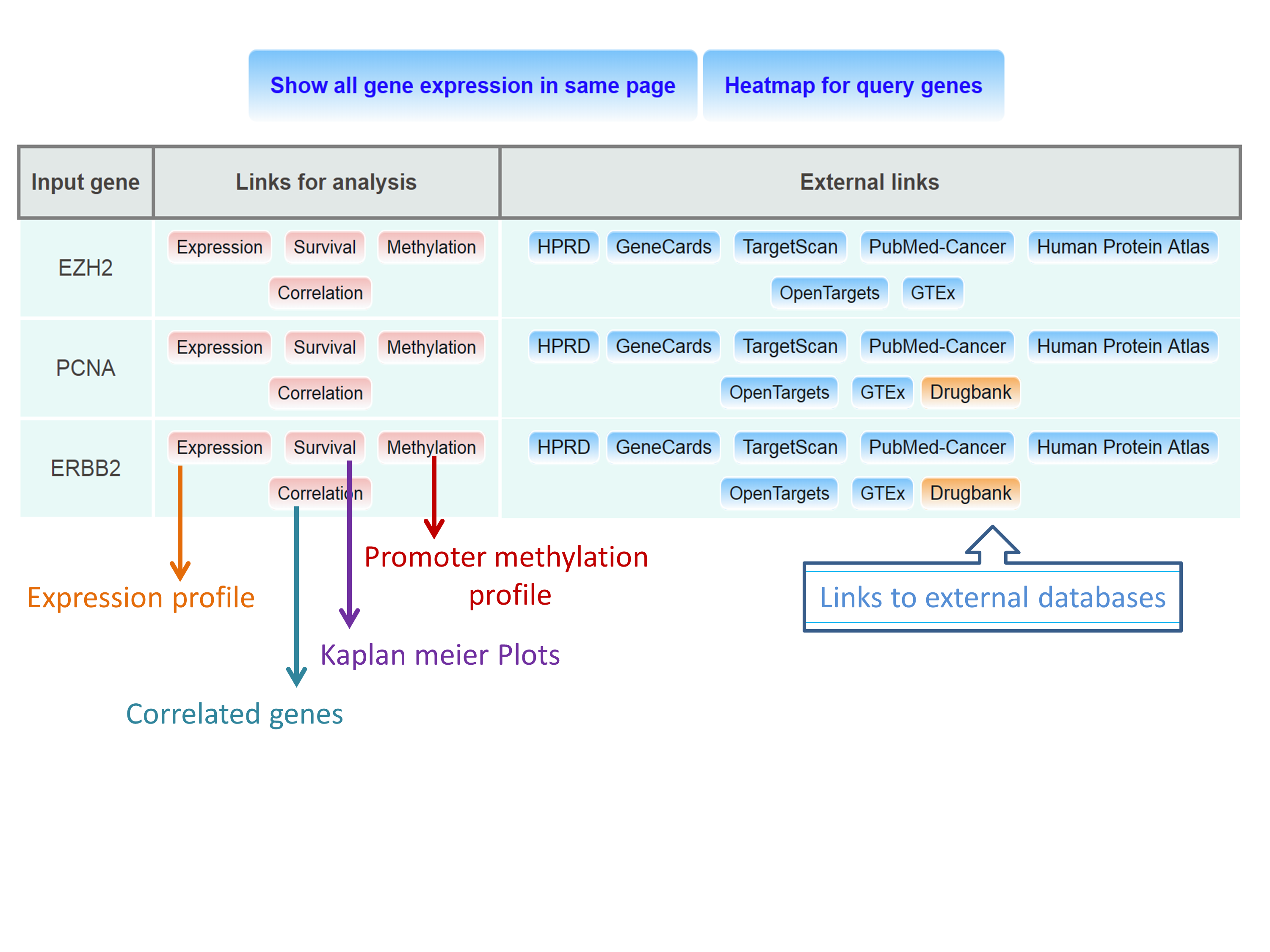
Step 3: Output page provides links to analysis results and external database links.
Obtaining EZH2 expression profile in breast invasive carcinoma dataset based on patient’s race.
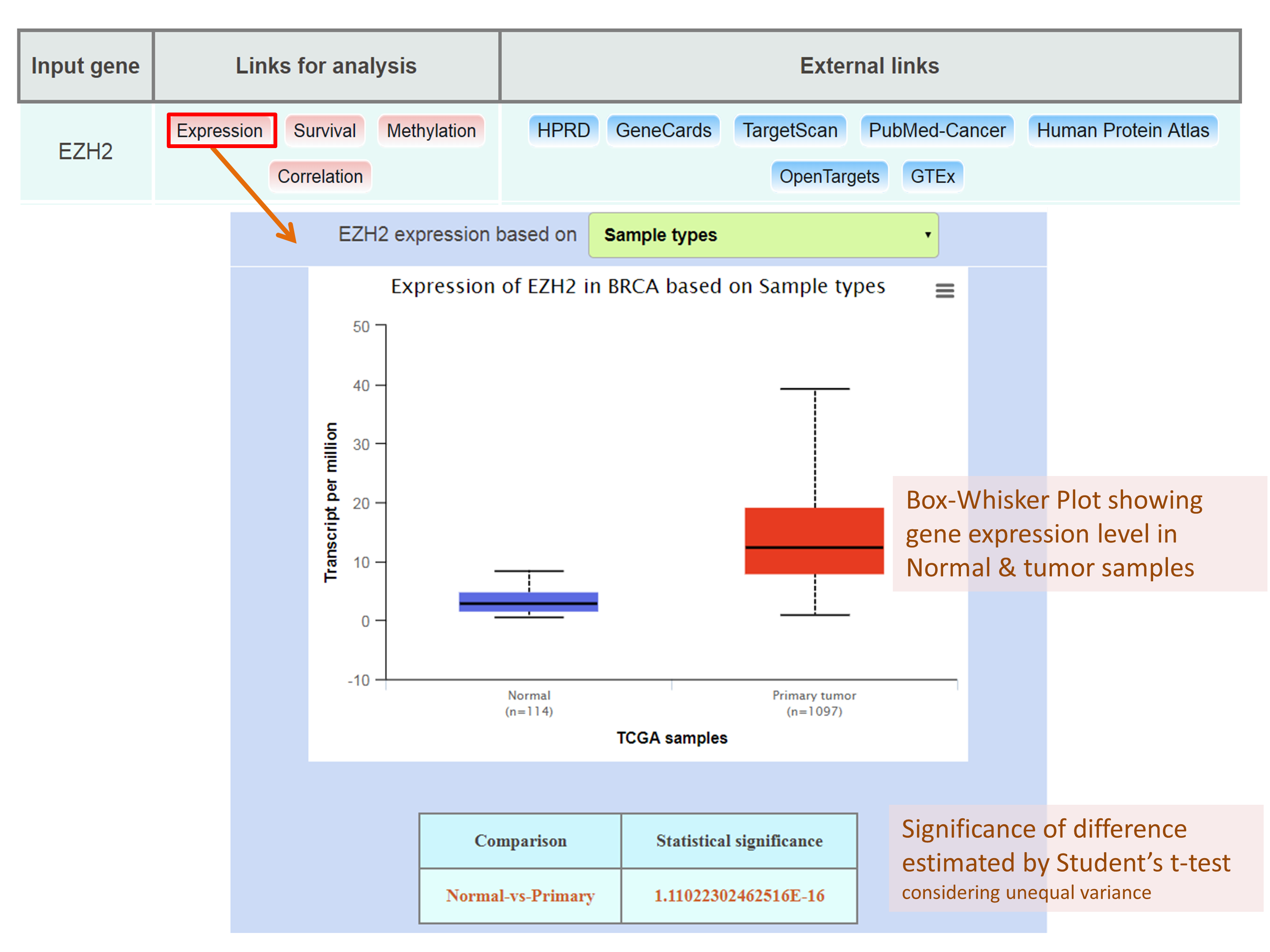
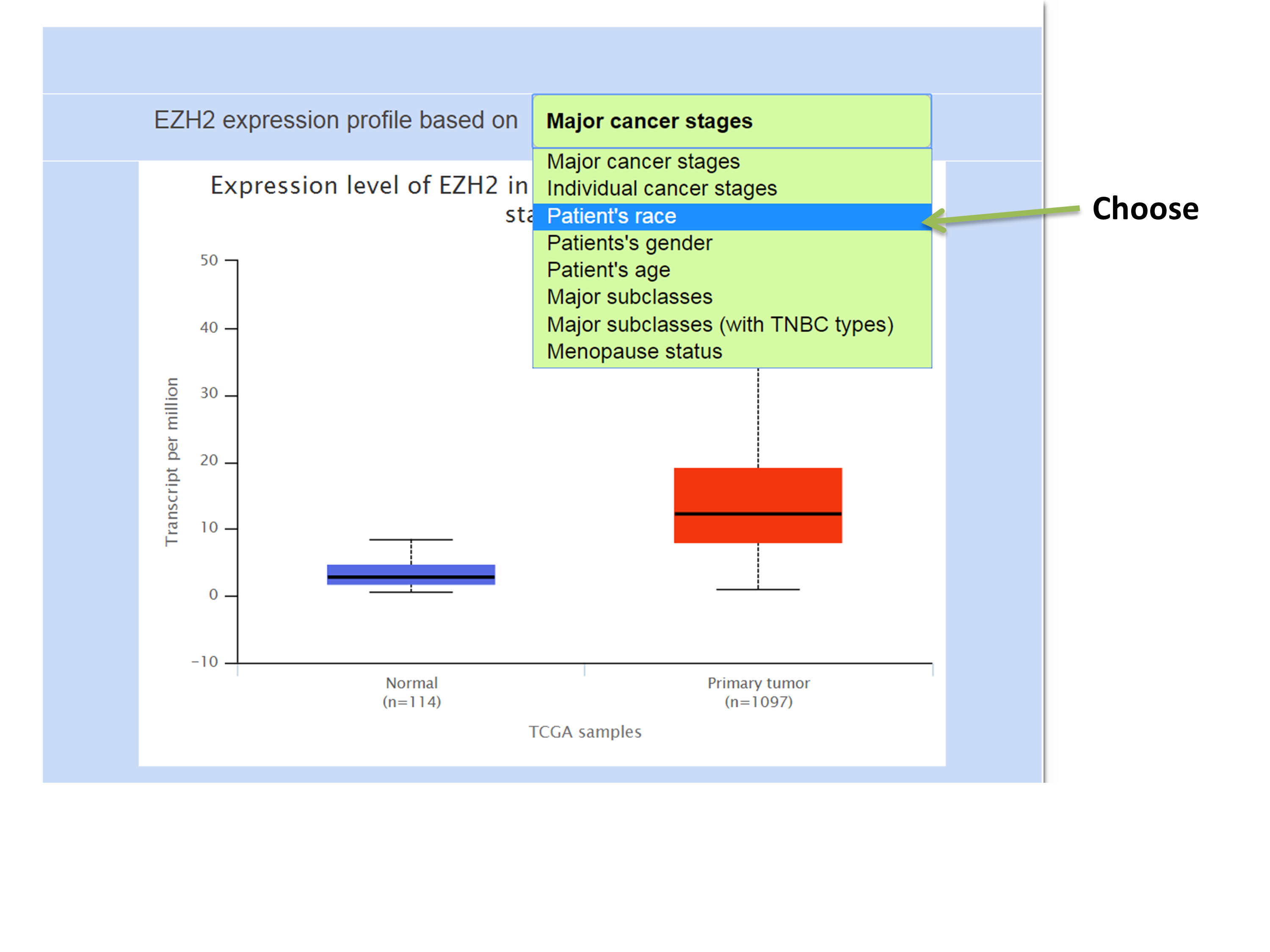
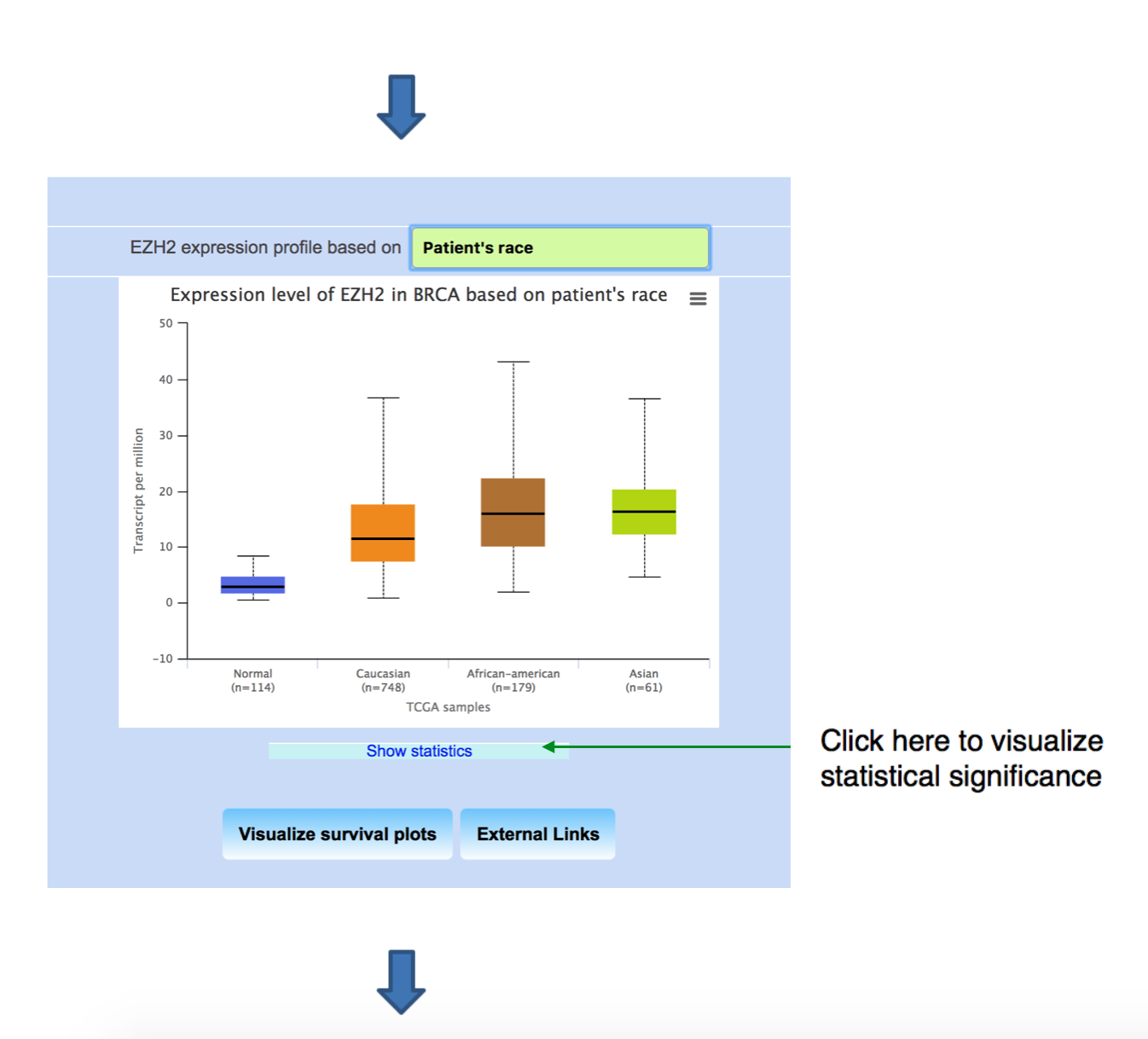
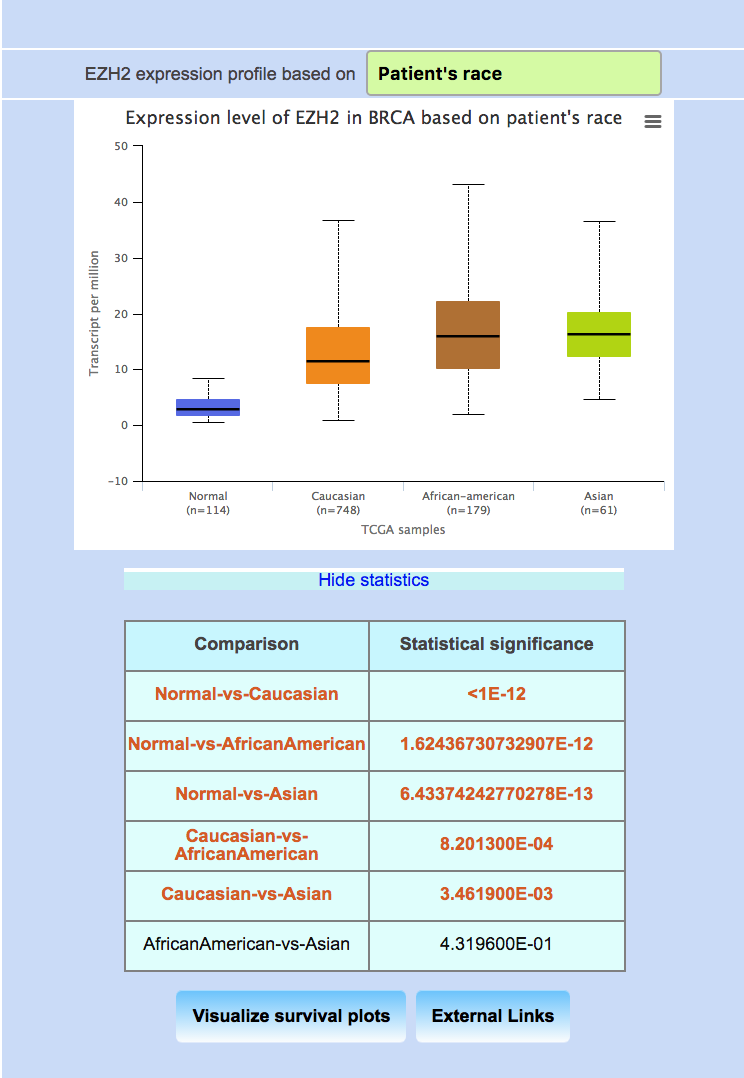
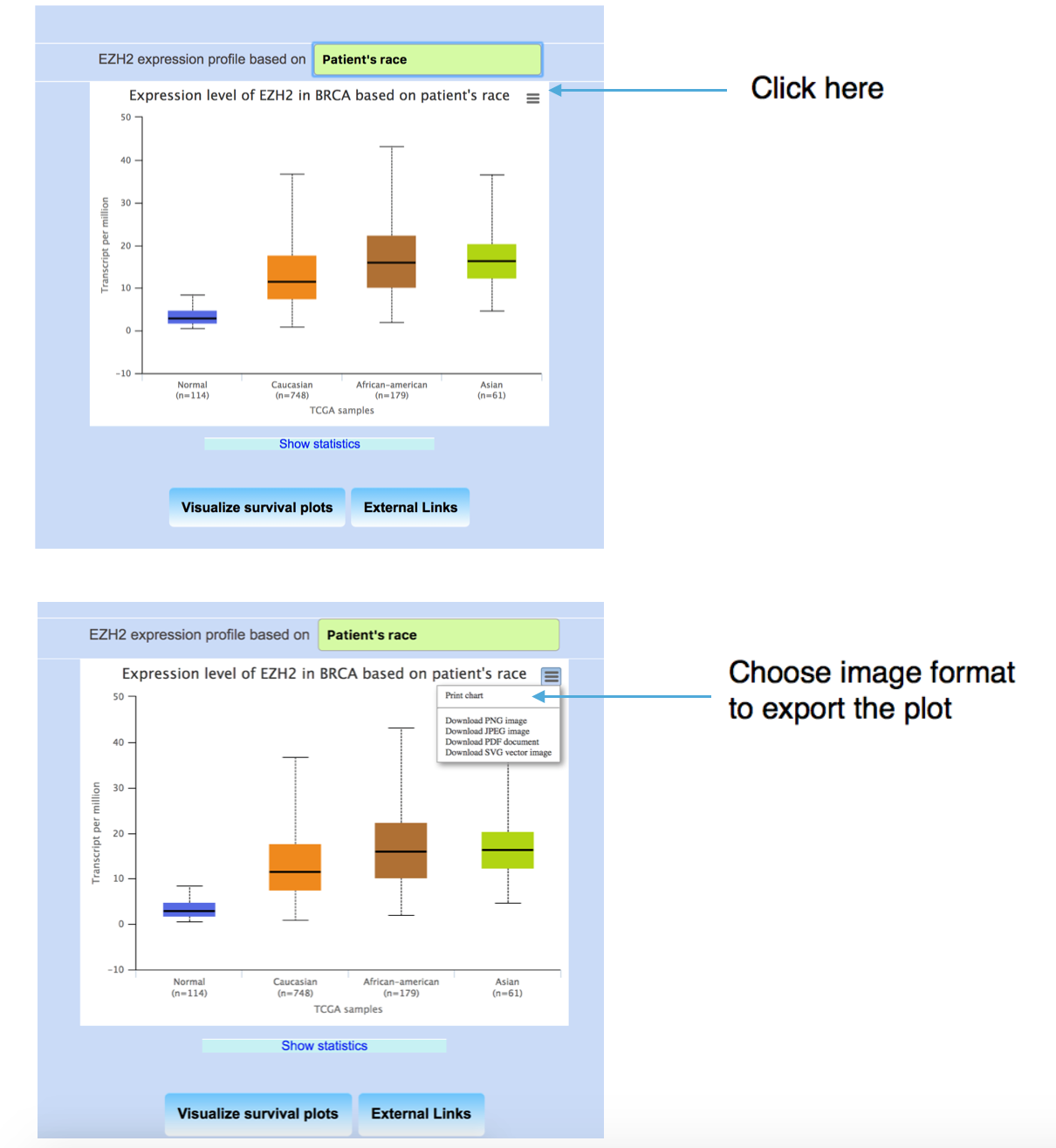
Downloading the box plot from UALCAN
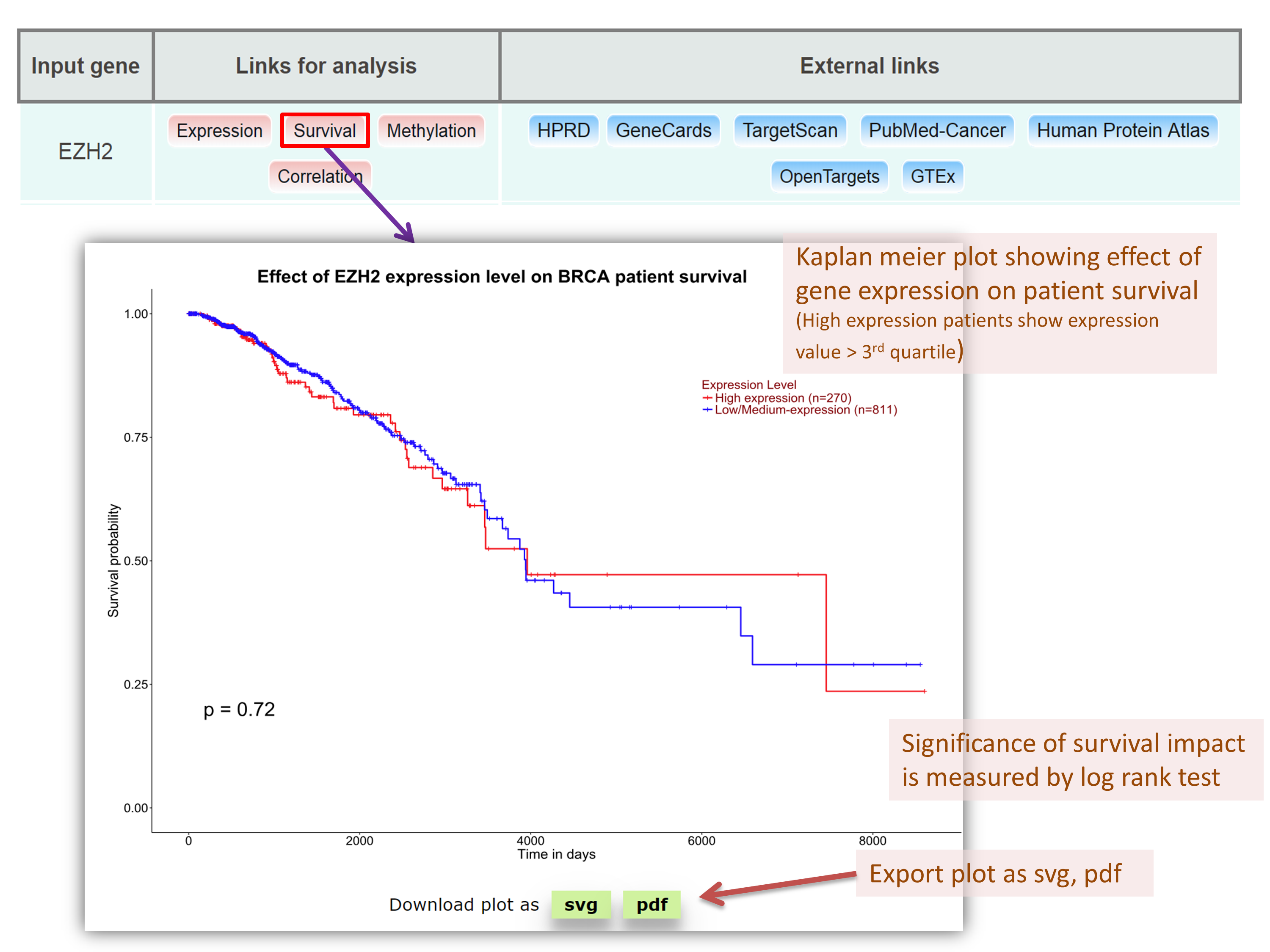
Analyzing survival plots of EZH2 in breast invasive carcinoma TCGA dataset
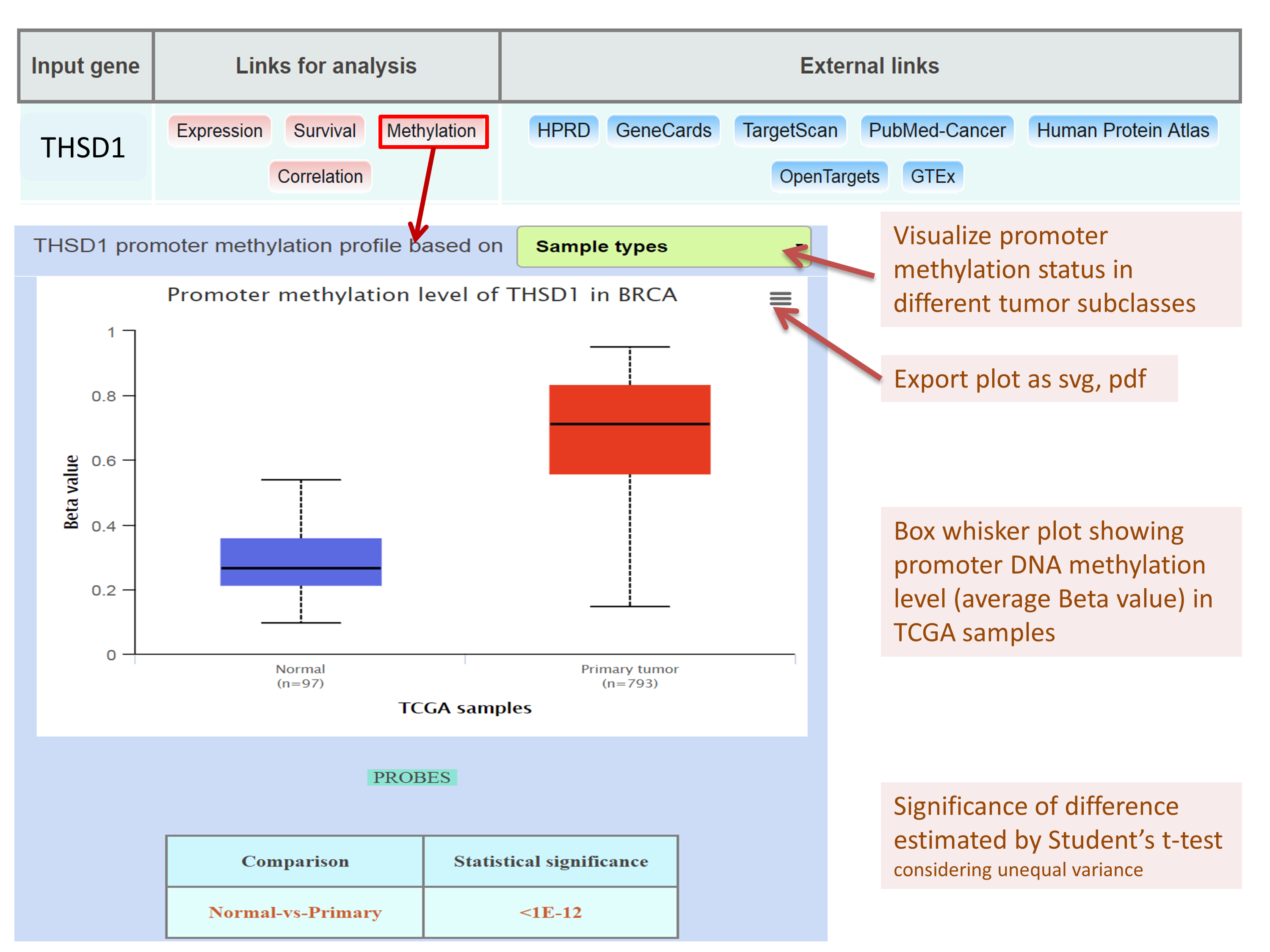
Analyzing promoter DNA methylation level of THSD1 in breast invasive carcinoma TCGA dataset
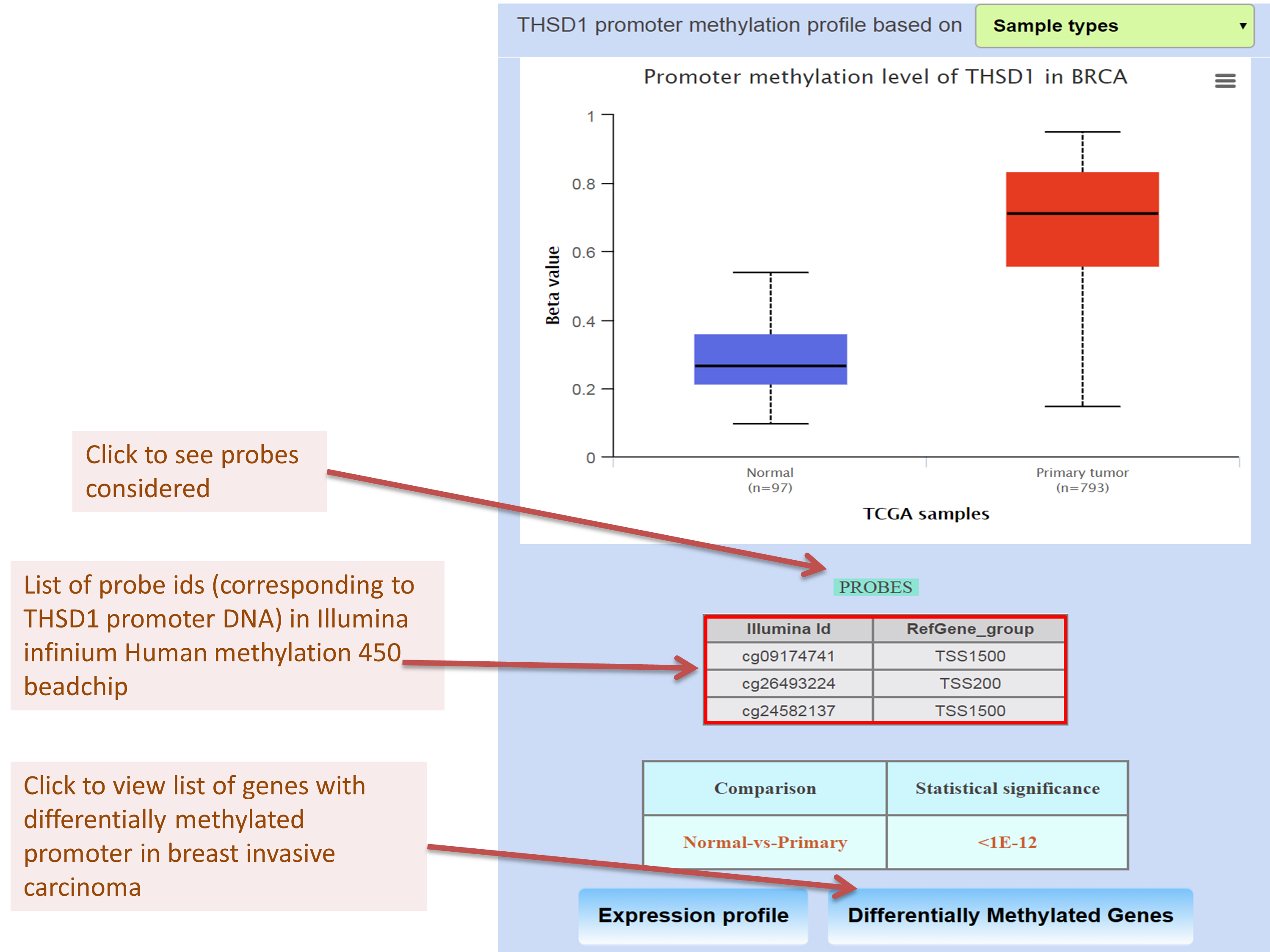
Obtaining list of positively/negatively correlated genes of EZH2 in breast invasive carcinoma TCGA dataset
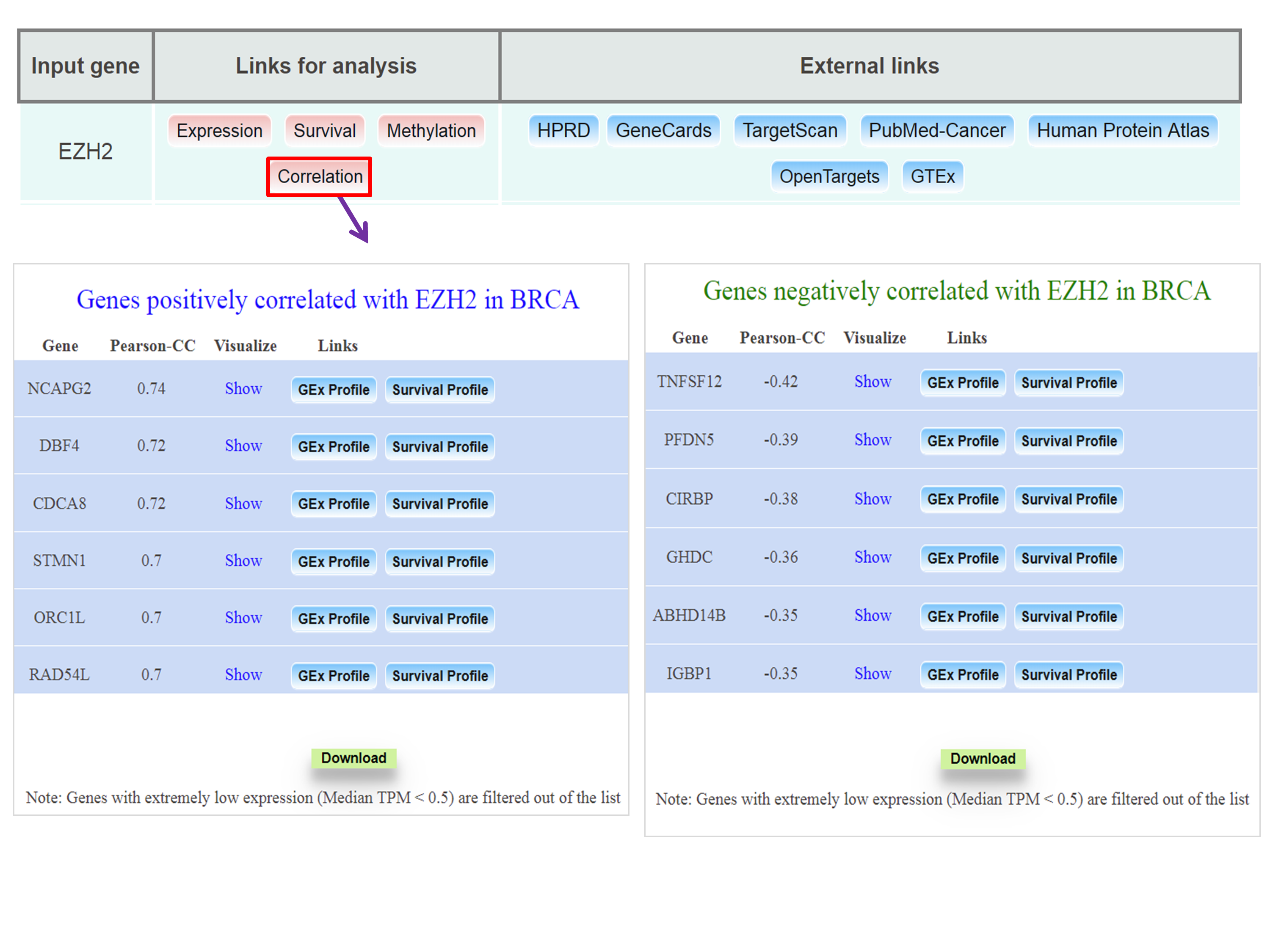
Step 1: Go to analysis page of UALCAN
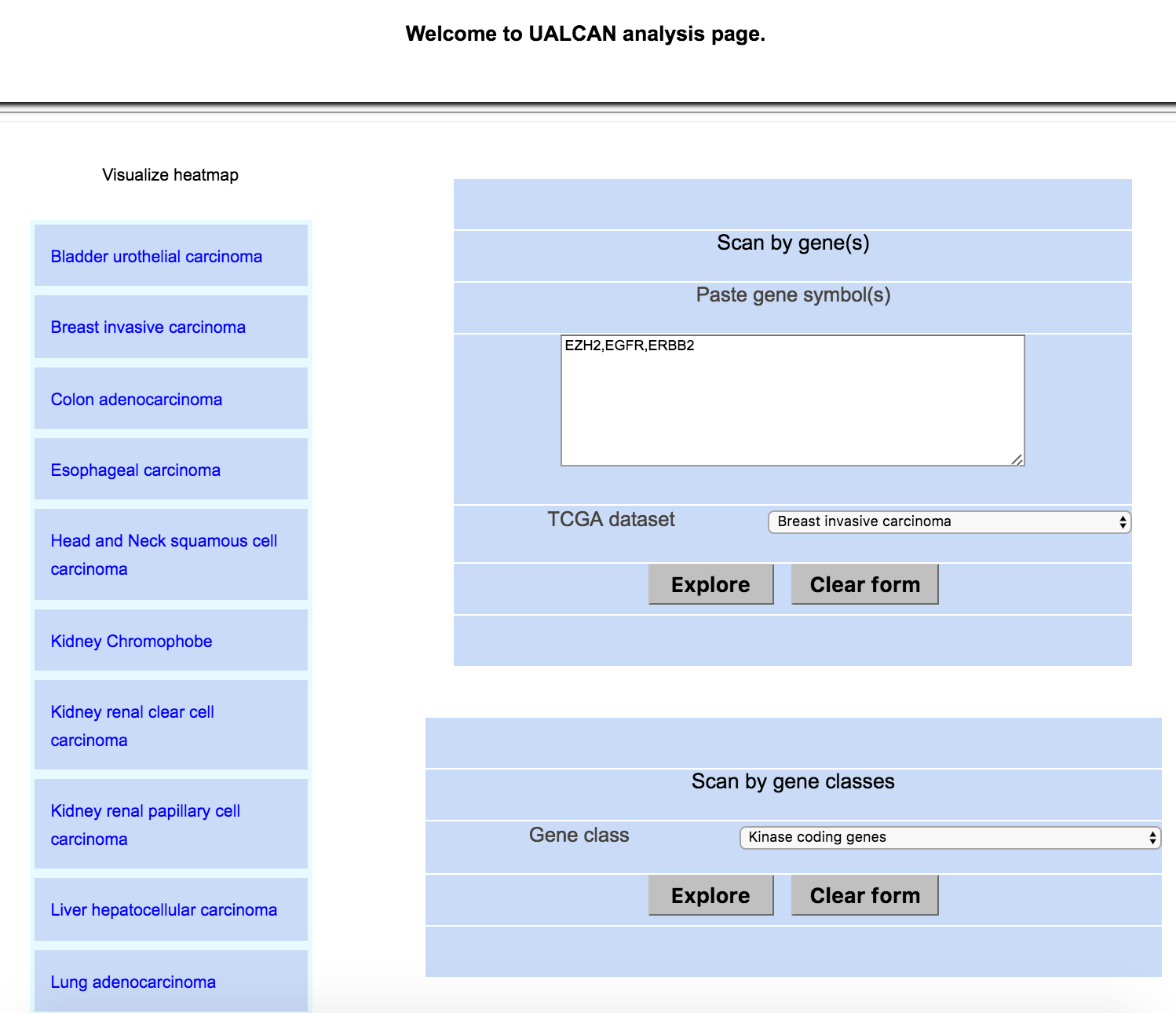

Step 2: Query UALCAN for pre-compiled gene classes
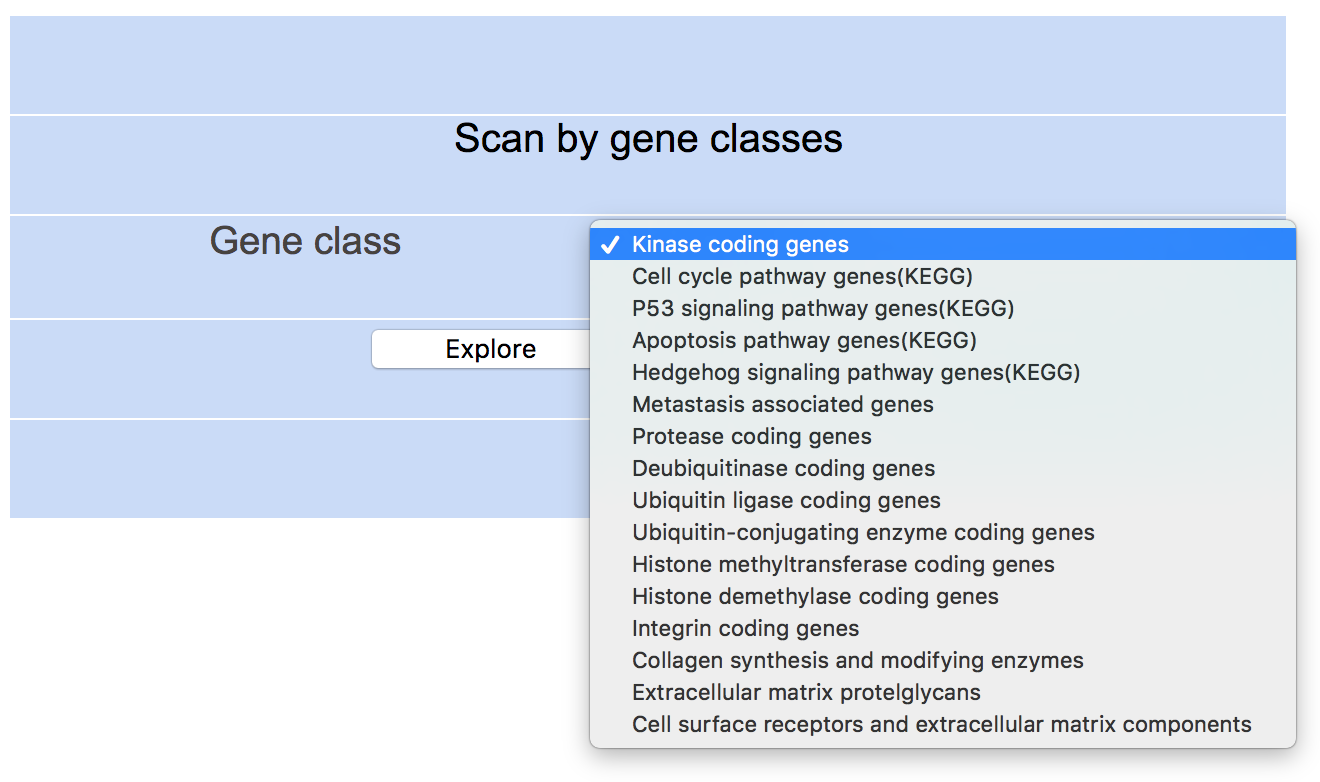
Step 3: Select gene class of interest (e.g. Kinase coding genes) and click "Explore" to see results
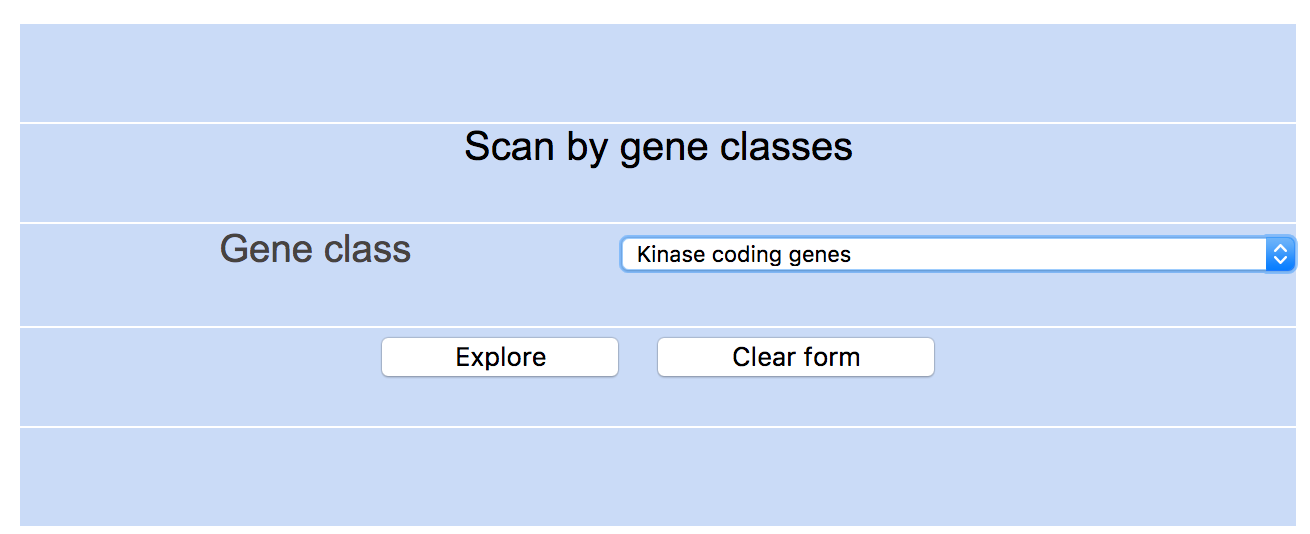
Step 4: UALCAN lists all kinase coding genes and indicates their expression status and survival effect in different cancers.
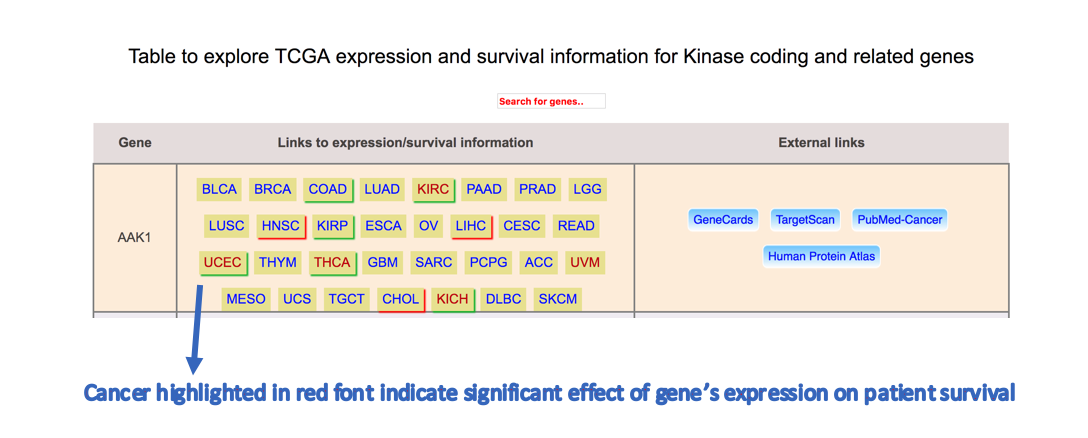
Cancer with red border show up regulation of gene
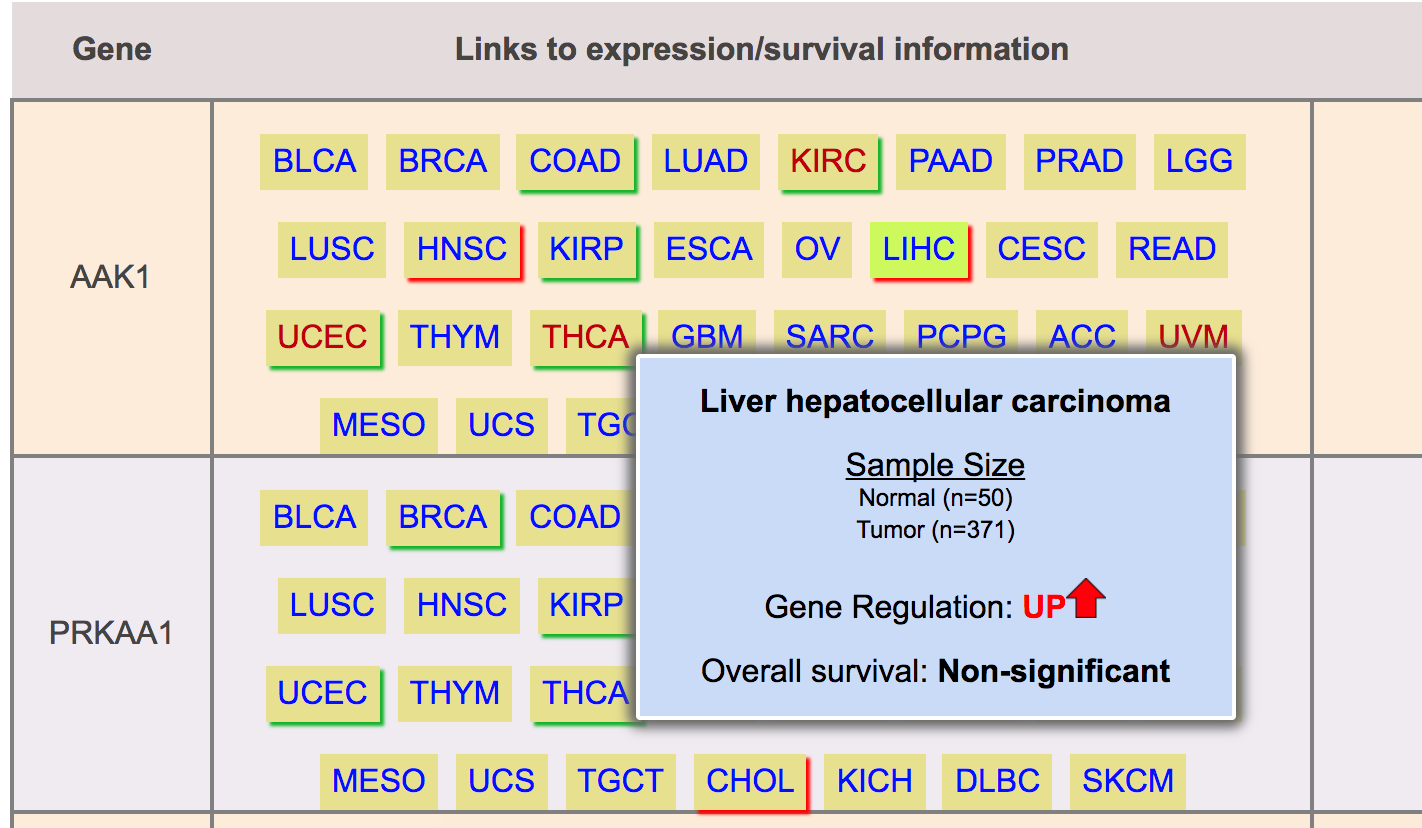
Cancer with green border show down regulation of gene
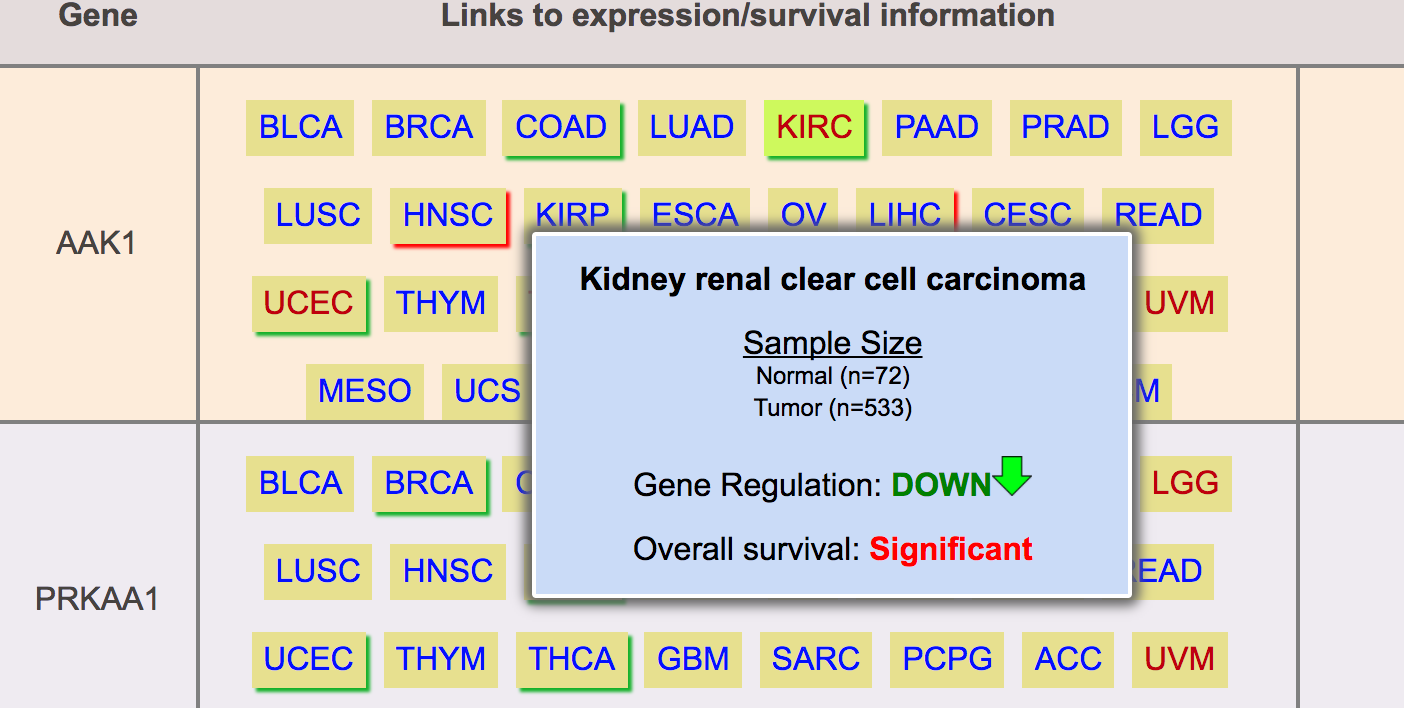
Step 5: User can click on any of these link to visualize expression and survival profiles
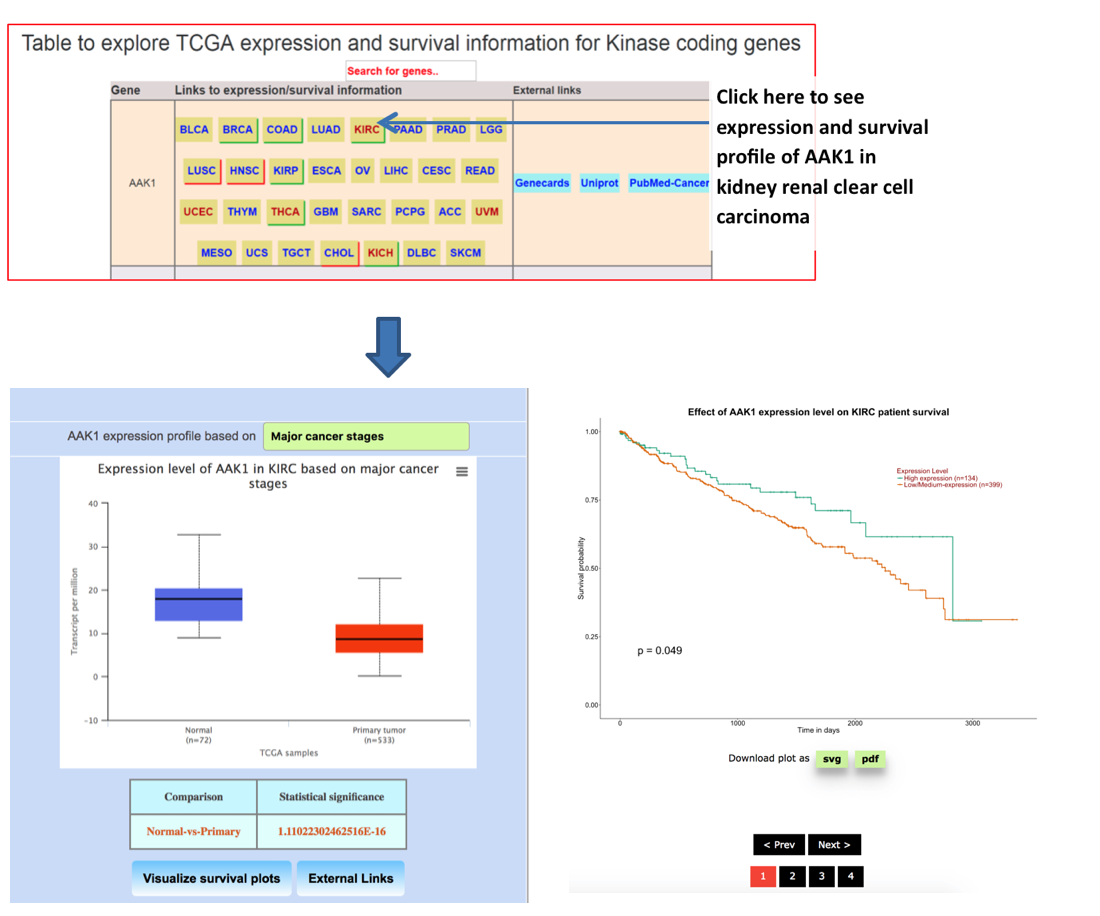
| Cancer | TCGA code |
| Adrenocortical carcinoma | ACC |
| Bladder urothelial carcinoma | BLCA |
| Brain lower grade glioma | LGG |
| Breast invasive carcinoma | BRCA |
| Cervical squamous cell carcinoma | CESC |
| Cholangiocarcinoma | CHOL |
| Colon adenocarcinoma | COAD |
| Esophageal carcinoma | ESCA |
| Glioblastoma multiforme | GBM |
| Head and Neck squamous cell carcinoma | HNSC |
| Kidney Choromophobe | KICH |
| Kidney renal clear cell carcinoma | KIRC |
| Kidney renal papillary cell carcinoma | KIRP |
| Liver hepatocellular carcinoma | LIHC |
| Lung adenocarcinoma | LUAD |
| Lung squamous cell carcinoma | LUSC |
| Lymphoid Neoplasm Diffuse Large B-cell Lymphoma | DBLC |
| Mesothelioma | MESO |
| Ovarian serous cystadenocarcinoma | OV |
| Pancreatic adenocarcinoma | PAAD |
| Pheochromocytoma and Paraganglioma | PCPG |
| Prostate adenocarcinoma | PRAD |
| Rectum adenocarcinoma | READ |
| Sarcoma | SARC |
| Skin Cutaneous Melanoma | SKCM |
| Testicular Germ Cell Tumors | TGCT |
| Thymoma | THYM |
| Thyroid carcinoma | THCA |
| Uterine Carcinosarcoma | UCS |
| Uterine Corpus Endometrial Carcinoma | UCEC |
| Uveal Melanoma | UVM |